This topic takes on average 55 minutes to read.
There are a number of interactive features in this resource:
 Chemistry
Chemistry
With traditional PCR, the amount of product can only be worked out at the end of the reaction (the plateau phase), typically by agarose gel electrophoresis. Conversely, real-time PCR (RT-PCR) allows for the simultaneous amplification and quantification of a targeted DNA molecule during a reaction. The procedure is essentially the same as the standard PCR method but amplified DNA is detected during the reaction as it progresses; in real time. A labelled probe is included in the PCR reagent mix in addition to the primers used in a traditional PCR reaction.
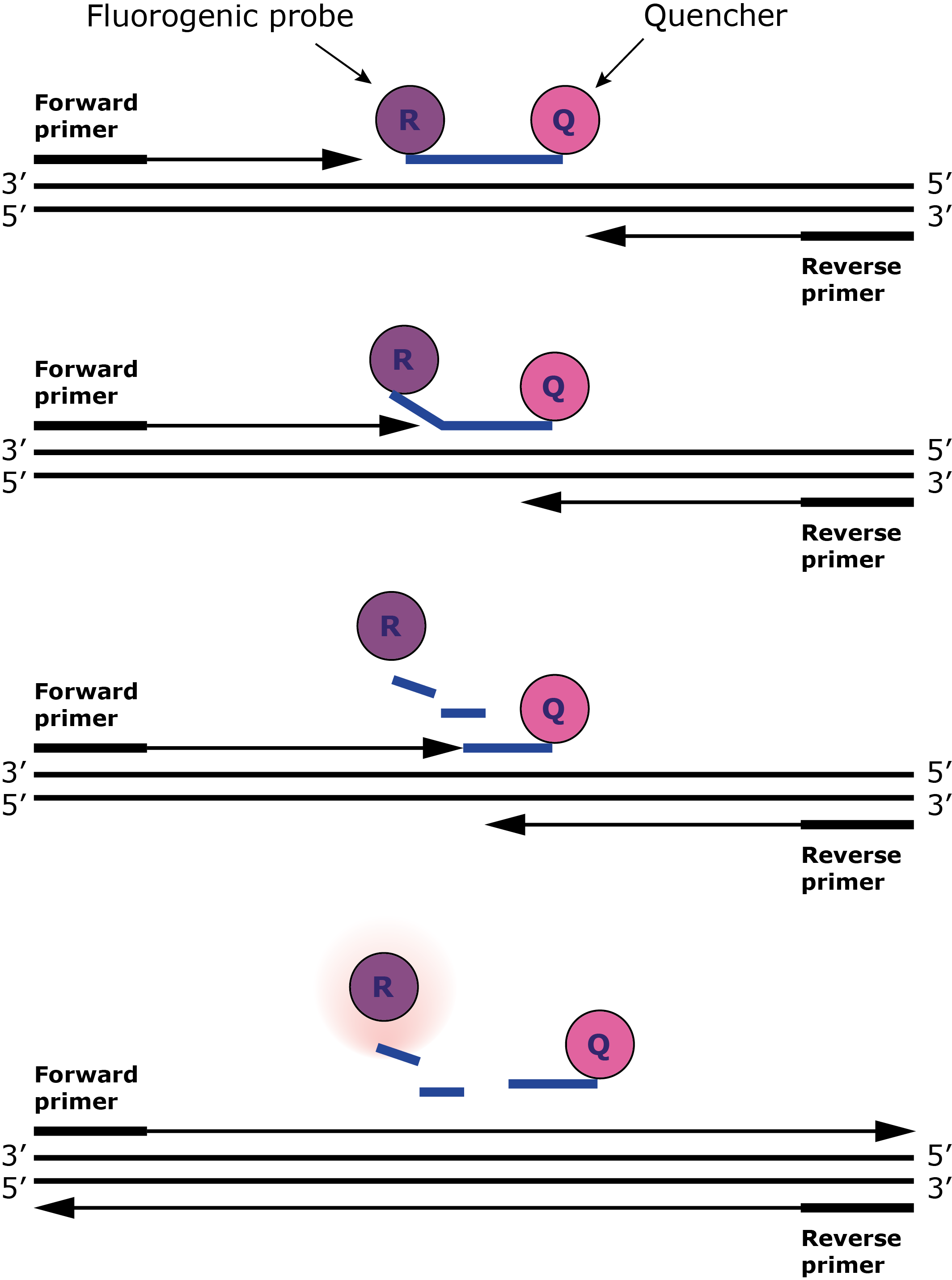
Sequence-specific DNA probes of oligonucleotides bind to the gene of interest. Oligonucleotides are short lengths of nucleic acid, usually with fewer than 50 nucleotides each comprising of a nitrogenous base, five-carbon sugar and one to three phosphate groups. They readily bind to lengths of DNA with the complementary sequence of bases (A with T, C with G) and therefore make excellent probes. In the case of RT-PCR, these are labelled with a fluorescent reporter dye at the 5‘ end and a quencher dye at the 3' end and only produce fluorescence when DNA polymerase moves along the template and cleaves (splits) the probe. Increased fluorescence intensity is coupled with an increase in the number of times the gene has been found in the sample.
The process:
|
RT- PCR is becoming ever more important in the diagnosis of disease strains – for example, in the outbreak of avian influenza which affected UK flocks in some areas in 2017. In an outbreak, it is vital to identify the strains and the sources of infection as rapidly as possible, to reduce the spread of the disease. Although the equipment for RT-PCR is expensive, the analysis itself is much faster – it can be completed in 3 hours if required – and cheaper than conventional PCR. Large numbers of samples can be tested, the process can identify both types and sub-types of the virus and scientists don’t need viable virus to make the identification. RT-PCR also plays an important role in the TB pathogen identification diagnostic system successfully introduced in the UK in 2017.
